NCN POLONEZ 2
dr Michał Szymański
Description
Oxidative stress leads to genomic instability and human diseases like cardiovascular, skeletal muscular and neurological disorders, cancer as well as normal aging process. Since mitochondrial DNA (mtDNA) is subjected to a constant attack by reactive oxygen species (ROS), generated as byproducts of oxidative phosphorylation, mitochondria must have robust DNA repair mechanisms. It is believed that base excision repair (BER) pathway is a major defense mechanism against oxidative damage in human mitochondria. A hallmark of BER is excision of 5′-DNA damage, most often 5′-deoxyribose phosphate (5′-dRP), from incised abasic site (AP site). If 5′-dRP is not removed it stalls the activity of DNA polymerase γ (Polγ), the only polymerase in human mitochondria. There are three candidates for the 5′-end damage removal in mitochondria: FEN1, DNA2 and EXOG. While FEN1 and DNA2 are shared by the nucleus and mitochondria, EXOG localizes only to mitochondria. Furthermore, depletion of EXOG causes accumulation of DNA damage in the mitochondria, but not in the nucleus, increases oxidative stress and mitochondrial dysfunction. EXOG has been recently shown to cleave a variety of different substrates; however, the molecular mechanism of its broadly defined activities, especially in the context of human mitochondrial BER, is not well understood. To provide fundamental mechanistic insights into human mitochondrial BER, we aim to examine the function of EXOG and Polγ on damaged DNA. Utilizing combination of biochemical and biophysical methods and building on recently established structural biology framework, we plan to dissect the role of these two crucial enzymes as they cooperate to repair human mtDNA damage.
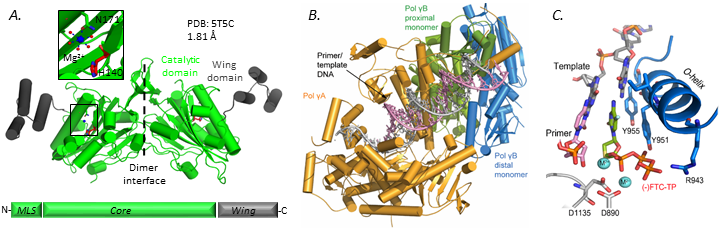
A. Crystal structure of EXOG (Szymanski, et al., Nature Communications 2017),
B. Polγ replicating complex (Szymanski et al., EMBO J 2015)
C. Polγ inhibited by (-)-FTC-TP, FDA approved antiviral drug for treatment of AIDS (Sohl* and Szymanski* et al., PNAS 2015)
Michał R. Szymański, PhD - Principal Investigator
Degrees:
2011 PhD in Biochemistry and Molecular, Biochemistry and Molecular Biology Department, Molecular Biophysics Educational Track, University of Texas Medical Branch at Galveston, Texas, USA (Supervisor: Prof. Wlodek M. Bujalowski)
2007 BSc in Biochemical and Biophysical Sciences, Biology and Biochemistry Department, University of Houston, Houston, Texas, USA
Research experience:
2017 - Present : Assistant Professor
2016 - 2017 Research Scientist in laboratory of Prof. Whitney Yin, Pharmacology and Toxicology Department, The University of Texas Medical Branch at Galveston, Texas, USA
2013 - 2016 Jeane B. Kempner Postdoctoral Fellow in laboratory of Prof. Whitney Yin, Pharmacology and Toxicology Department, The University of Texas Medical Branch at Galveston, Texas, USA
2012 - 2013 Jeane B. Kempner Postdoctoral Fellow in laboratory of Prof. Wlodek M. Bujalowski, Biochemistry and Molecular Biology, The University of Texas Medical Branch at Galveston, Texas, USA
2007 - 2011 Graduate Assistant in laboratory of Prof. Wlodek M. Bujalowski, Biochemistry and Molecular Biology, The University of Texas Medical Branch at Galveston, Texas, USA
2005 - 2007 Welch Research Assistant in laboratory of Prof. H.J. Yeo, Biology and Biochemistry Department, University of Houston, Houston, Texas, USA
2004 - 2005 Research Assistant, Polyorganix Inc., Houston, Texas, USA
Publications:
- Szymanski, M.R., Yu, A., Gmyrek, A.M., White, M.A., Molineux, I.J., Lee, J.C., Yin W.Y. A novel domain in human EXOG converts apoptotic endonuclease to DNA- repair exonuclease. Nature Communications. 2017 May 3;8:14959.
Recommended by F1000. - Li, M., Mislak, A.C., Foli, Y., Agbosu, E., Bose, V., Bhandari, S., Szymanski, M.R., Shumate, C.K., Yin, W., Anderson, K.S., Paintsil, E. DNA Polymerase-γ R953C Mutant Linked to ART-Associated Mitochondrial Toxicity. Antimicrobial Agents and Chemotherapy. 2016 Aug 22;60(9):5608-11.
- Sohl, C.D*., Szymanski, M.R*., Mislak, A.C., Shumate, K.C., Amiralaei, S., Schinazi, F.R., Anderson K.S., Yin W.Y. Probing the Structural and Molecular Basis of Nucleotide Selectivity by Human Mitochondrial DNA Polymerase γ. Proceedings of National Academy of Sciences of the United States of America. 2015 Jul 14;112(28):8596-601.
*equal contribution - Szymanski, M.R., Kuznetsov, V.B., Shumate, C., Meng, Q., Lee, Y-S., Patel, G., Patel, S.S., Yin W.Y. Structural basis for processivity and antiviral drug toxicity in human mitochondrial DNA replicase. EMBO J. 2015 Jul 14; 34(14):1959-70.
- Szymanski, M.R., Jezewska, M.J., Bujalowski, W. The Escherichia Coli Primosomal DnaT Protein Exists in Solution as a Monomer – Trimer Equilibrium System. Biochemistry. 2013 Mar 19;52(11):1845-57.
- Szymanski, M.R., Jezewska, M.J., Bujalowski, W. Energetics of the Escherichia Coli DnaT Protein Trimerization Reaction. Biochemistry. 2013 Mar 19;52(11):1858-73.
- Szymanski, M.R., Bujalowski, P.J., Jezewska, M.J., Gmyrek, A.M., and Bujalowski, W. The N-terminal domain of the Escherichia coli PriA helicase contains both the DNA- and nucleotide-binding sites. Energetics of domain-DNA interactions and allosteric effect of the nucleotide cofactors. Biochemistry. 2011 Nov 1;50(43):9167-83.
- Szymanski, M.R., Jezewska, M.J., and Bujalowski, W. Binding of Two PriAPriB Complexes to the Primosome Assembly Site Initiates Primosome Formation. Journal of Molecular Biology. 2011 Aug 5;411(1):123-42.
- Szymanski, M.R., Jezewska, M.J., Bujalowski, P.J., Bussetta, C., Ye, M., Choi K.H., Bujalowski, W. Full-Length Dengue Virus RNA Dependent RNA Polymerase – RNA/ DNA Complexes. Stoichiometries and Energetics of Intrinsic Affinities, Cooperativities, Base and Conformational Specificities. Journal of Biological Chemistry. 2011 Sep 23;286(38):33095-108.
- Jezewska, M.J., Szymanski, M.R., and Bujalowski, W. Kinetic mechanism of the ssDNA recognition by the polymerase X from African Swine Fever Virus. Dynamics and energetics of intermediate formations. Biophysical Chemistry. 2011 Sep;158(1):9-20.
- Jezewska, M.J., Szymanski, M.R., and Bujalowski, W. Interactions of the DNA polymerase X from African Swine Fever Virus with the ssDNA. Properties of the total DNA-binding site and the strong DNA-binding subsite. Biophysical Chemistry. 2011 Sep;158(1):26-37.
- Jezewska, M.J., Szymanski, M.R., and Bujalowski, W. The primary DNAbinding subsite of the rat pol beta. Energetics of interactions of the 8-kDa domain of the enzyme with the ssDNA. Biophysical Chemistry. 2011 Jul;156(2-3):115-27.
- Szymanski, M.R., Jezewska, M.J., and Bujalowski, W. The Escherichia coli PriA helicase-double-stranded DNA complex: location of the strong DNA-binding subsite on the helicase domain of the protein and the affinity control by the two nucleotide-binding sites of the enzyme. Journal of Molecular Biology. 2010 Sep 17;402(2):344-62.
- Szymanski, M.R., Jezewska, M.J., and Bujalowski, W. Interactions of the Escherichia coli primosomal PriB protein with the single-stranded DNA. Stoichiometries, intrinsic affinities, cooperativities, and base specificities. Journal of Molecular Biology. 2010 Apr 23;398(1):8-25.
- Szymanski, M.R., Jezewska, M.J., and Bujalowski, W. The Escherichia coli PriA helicase specifically recognizes gapped DNA substrates: effect of the two nucleotide-binding sites of the enzyme on the recognition process. Journal of Biological Chemistry. 2010 Mar 26;285(13):9683-96.
- Andreeva, I.E., Roychowdhury, A., Szymanski, M.R., Jezewska, M.J., and Bujalowski, W. Mechanisms of interactions of the nucleotide cofactor with the RepA protein of plasmid RSF1010. Binding dynamics studied using the fluorescence stopped-flow method. Biochemistry. 2009 Nov 10;48(44):10620-36.
- Szymanski, M.R., Fiebach, A.R., Tratschin, J.D., Gut, M., Ramanujam, V.M., Gottipati, K., Patel, P., Ye, M., Ruggli, N., and Choi, K.H. Zinc binding in pestivirus N(pro) is required for interferon regulatory factor 3 interaction and degradation. Journal of Molecular Biology. 2009 Aug 14;391(2):438-49.
- Roychowdhury, A., Szymanski, M.R., Jezewska, M.J., and Bujalowski, W. Interactions of the Escherichia coli DnaB-DnaC protein complex with nucleotide cofactors. 1. Allosteric conformational transitions of the complex. Biochemistry. 2009 Jul 28;48(29):6712-29.
- Roychowdhury, A., Szymanski, M.R., Jezewska, M.J., and Bujalowski, W. Mechanism of NTP hydrolysis by the Escherichia coli primary replicative helicase DnaB protein. 2. Nucleotide and nucleic acid specificities. Biochemistry. 2009 Jul 28;48(29):6730-46.
- Roychowdhury, A., Szymanski, M.R., Jezewska, M.J., and Bujalowski, W. Escherichia coli DnaB helicase-DnaC protein complex: allosteric effects of the nucleotides on the nucleic acid binding and the kinetic mechanism of NTP hydrolysis. 3. Biochemistry. 2009 Jul 28;48(29):6747-63.
- Andreeva, I.E., Szymanski, M.R., Jezewska, M.J., Galletto, R., and Bujalowski, W. Dynamics of the ssDNA recognition by the RepA hexameric helicase of plasmid RSF1010: analyses using fluorescence stopped-flow intensity and anisotropy methods. Journal of Molecular Biology. 2009 May 15;388(4):751-75.
- Duret, G., Szymanski, M., Choi, K.J., Yeo, H.J., and Delcour, A.H. The TpsB translocator HMW1B of haemophilus influenzae forms a large conductance channel. Journal of Biological Chemistry. 2008 Jun 6;283(23):15771-8.
Financing
 This project has received funding from the European Union’s Horizon 2020 research and innovation programme under the Marie Skłodowska-Curie Actions.
This project has received funding from the European Union’s Horizon 2020 research and innovation programme under the Marie Skłodowska-Curie Actions.
Grant agreement No. 665778.
 Project is conducted within POLONEZ funding programme of National Science Centre POLAND.
Project is conducted within POLONEZ funding programme of National Science Centre POLAND.
Fellowship registration number: 2016/21/P/NZ1/01085





