NCN POLONEZ BIS 2
Dynamics and evolution of specificity of TA complexes
TA dynamics
Toxin-antitoxin (TA) systems are ubiquitous genetic modules promoting bacterial survival by regulation and arrest of cell growth in response to stresses such as phage infection, antibiotic exposure, or macrophage uptake. While toxins target essential biological processes affecting growth of a cell, antitoxins specifically neutralize their cognate toxins and allow for cell growth. In the most prevalent type II TAs, the antitoxin is a protein that directly binds its cognate toxin with a neutralization domain (ND), resulting in formation of a tight toxin-antitoxin complex. Besides the ND, the antitoxin also contains a DNA binding domain enabling it to transcriptionally control the toxin-antitoxin operon. Another key feature of TAs is the common presence of several paralogues of the same system within the same organism. The frequent complete insulation of numerous paralogous TAs, where there is high specificity with no cross-pairing, suggests that a neutralizing cross-interaction is detrimental to the cell. We are interested in studying the evolution of specificity of paralogous TA systems through toxin size variation by addition of structural add-on elements then allowing the antitoxin to evolve a specific neutralizing sequence. We found that such size alteration occurs at either N- or C-terminal toxin ends in various TA systems. Building on the experience gained while studying three paralogous Salmonella TacAT1-3 TA systems, we will continue to explore the significance of add-ons in contribution to development of neutralization specificity reducing paralogue crosstalk conflict. We will also investigate the activation of an important family of TAs involved in persistence of pathogenic Salmonella. Overall, these studies will provide new insights into the dynamics of evolution of paralogous protein-protein complexes and protein-DNA complexes, thereby opening avenues of interfering with these important stress response elements in bacteria.
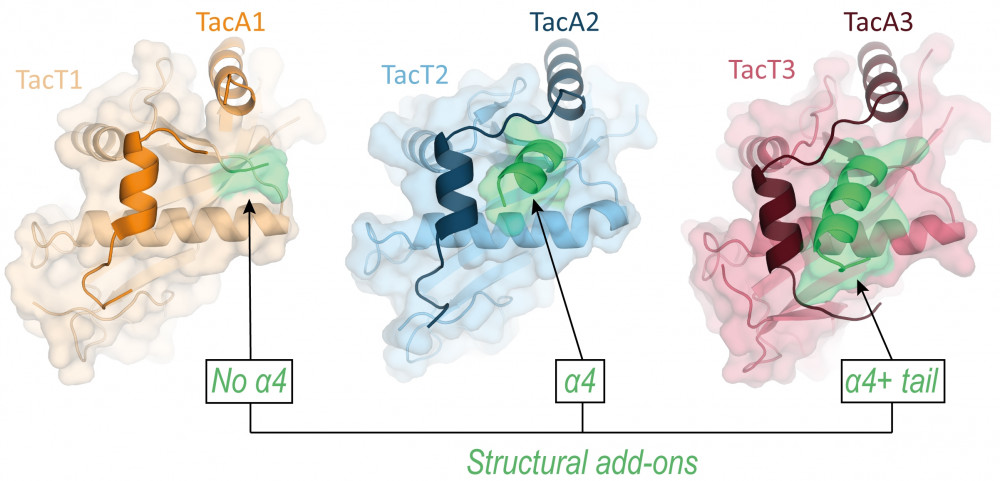
Structures of S. enterica TacT1-3 toxins neutralized by their cognate TacA1-3 antitoxins.
Green color points to the variable TacT C-terminal add-on.
Grzegorz J. Grabe, PhD – Principal Investigator
Structural Biology Laboratory
email: grzegorz.grabe@ug.edu.pl
phone: +48 58 523 6434
Degrees:
2016 – PhD, Imperial College London, UK
2011 – MRes, Imperial College London, UK
2010 – MSc, University of Gdansk and Medical University of Gdansk, Poland
2007 – BSc, University of Gdansk and Medical University of Gdansk, Poland
Selected publications:
- Grabe GJ, Giorgio RT, Wieczór M, Gollan B, Sargen M, Orozco M, Hare SA, Helaine S. Molecular stripping underpins derepression of a toxin–antitoxin system. Nat Struct Mol Biol 2024
- Worm DJ, Grabe GJ, de Castro GV, Rabinovich S, Warm I, Isherwood K, Helaine S, and Barnard A. Stapled Phd Peptides Inhibit Doc Toxin Induced Growth Arrest in Salmonella. ACS Chem Biol 2023
- De Castro GV, Worm DJ, Grabe GJ, Rowan FC, Haggerty L, De La Lastra AL, Popescu O, Helaine S & Barnard A. Characterization of the Key Determinants of Phd Antitoxin Mediated Doc Toxin Inactivation in Salmonella. ACS Chem Biol 2022
- Grabe GJ, Giorgio RT, Hall AMJ, Morgan RML, Dubois L, Sisley TA, Rycroft JA, Hare SA, Helaine S. Auxiliary interfaces support the evolution of specific toxin–antitoxin pairing. Nat Chem Biol, 2021
- Matthews-Palmer TRS, Gonzalez-Rodriguez N, Calcraft T, Lagercrantz S, Zachs T, Yu XJ, Grabe GJ, Holden DW, Nans A, Rosenthal PB, et al. Structure of the cytoplasmic domain of SctV (SsaV) from the Salmonella SPI-2 injectisome and implications for a pH sensing mechanism. J Struct Biol, 2021
- Rycroft JA, Gollan B, Grabe GJ, Hall AMJ, Cheverton AM, Larrouy-Maumus G, Hare SA, Helaine S. Activity of acetyltransferase toxins involved in Salmonella persister formation during macrophage infection. Nature Commun, 2018
- Yu X-J, Grabe GJ, Liu M, Mota LJ & Holden DW. SsaV interacts with SsaL to control the translocon-to-effector switch in the Salmonella SPI-2 type three secretion system. MBio, 2018
- Grabe GJ, Zhang Y, Przydacz M, Rolhion N, Yang Y, Pruneda JN, Komander D, Holden DW & Hare SA. The Salmonella effector SpvD is a cysteine hydrolase with a serovar-specific polymorphism influencing catalytic activity, suppression of immune responses, and bacterial virulence. J Biol Chem, 2016
- Rolhion N, Furniss RCD, Grabe GJ, Ryan A, Liu M, Matthews SA & Holden DW. Inhibition of Nuclear Transport of NF-ĸB p65 by the Salmonella Type III Secretion System Effector SpvD. PLoS Pathog, 2016
Job offer:
Research assistant (Deadline for submitting offers: June 30, 2023, 23:59) (outdated)
Results of the competition
This research is performed in a Structural Biology Group lead by Prof. Michal Szymanski. The group studies large macromolecular machines involved in DNA replication and repair processes and is located at Intercollegiate Faculty of Biotechnology in Gdansk.
Financing
This research is part of the project No. 2022/45/P/NZ1/03666 co-funded by the National Science Centre and the European Union’s Horizon 2020 research and innovation programme under the Marie Skłodowska-Curie grant agreement No 945339.

News:
-
I am excited to announce that I have been awarded an EMBO Installation Grant! This funding will enable me to establish an independent research group, where we will investigate Salmonella survival and pathogenesis. I am grateful for the support from EMBO and look forward to this new chapter. https://www.embo.org/press-releases/ten-group-leaders-awarded-embo-installation-grants/
-
This Wednesday (18.12.2024) I was thrilled to deliver a talk as a part of the Gdańsk University of the Third Age seminar series. We discussed Salmonella pathogenesis, persister cell phenomenon, and toxin-antitoxin stress response systems. The opportunity to share my knowledge and spark curiosity among the audience was indeed a rewarding experience

-
I had a great time in Kraków last week at the 26th Heart of Europe Bio-Crystallography Meeting! It was a fantastic opportunity to connect with brilliant structural biologists from across Central Europe and dive into their cutting-edge research. I also had the chance to share some of my latest findings. Huge thanks to the amazing organizers at the Małopolska Centre of Biotechnology, Jagiellonian University in Kraków, Poland!

-
In the past two weeks, I had the privilege of attending two prestigious scientific conferences: the 9th Central European Congress of Life Sciences Eurobiotech in Kraków, Poland (June 27-28, 2024) and the 48th FEBS Congress entitled "Mining Biochemistry for Human Health" in Milan, Italy (June 29 - July 3, 2024).
In Kraków, I engaged with leading researchers from Jagiellonian University and the Polish Academy of Science. I also gained valuable insights into the current state of the Polish biotechnology sector through discussions with representatives from prominent companies such as Molecure, Ryvu Therapeutics, Captor Therapeutics, and JJP Biologics.
In Milan, I had the honor of attending seminars led by world-renowned scientists and Nobel Laureates, including Bruce Beutler, Robert Huber, Venki Ramakrishnan, and John Walker. Over the four days of the Congress, I participated in numerous lectures covering a wide range of topics, such as advancements in structural biology, translational proteomics, novel antibacterial compound screens, and therapies focused on targeting cancer metabolism.
These past two weeks have been immensely stimulating, significantly expanding my network within the scientific community.
- I am thrilled to announce that I have been awarded an OPUS-26 grant from the Polish National Science Centre! This funding will enable us to further expand our research on the survival and virulence strategies of the Salmonella pathogen

- Over the last two weeks (8-18.04.2024), Magda and I have been learning about the “phage way” from super friendly and passionate scientists at Acteryon company. Bacteriophage-based treatments of recalcitrant bacterial infections have tremendous potential!

- On Wednesday (20.03.2024), IFB hosted the final stage of another round of the Biotechnological Knowledge Competition for high school students, with 'Basis of Tumorigenesis' as this year's main theme. I delivered a guest seminar in which we discussed Salmonella pathogenesis, treatment options, toxin-antitoxin systems, and how persister cells present therapeutic challenges in both bacterial infections and tumors. It was a pleasure to engage with the students and address their thought-provoking questions during the session!

- Last Friday (01.03.2024), I had the pleasure of presenting a seminar on toxin-antitoxin systems at the Institute of Microbiology of the Czech Academy of Sciences, located in Prague. During my stay, I met some amazing and driven scientists who work on Salmonella and Bordetella pathogens. We exchanged our experiences, shared ideas, and discussed potential collaboration.

- We are excited to host Dr Severin Ronneau from University of Namur (Belgium) at our faculty seminar series this Friday (26th Jan)! Severin will present his research on bacterial persistence in the context of infection. Link to the talk
- 30.11.-01.12.2023 – With my mentor, Michal Szymanski, we had an exciting kick-off meeting at the National Science Centre in Krakow. This was a fantastic opportunity to meet other Polonez BIS programme participants and their mentors. We also learnt a great deal about open access publishing and career development during dedicated workshops. It was an all-around fantastic experience






